IR Microscope Imaging Analysis Software
Spectra ManagerTM Micro Imaging Analysis Program
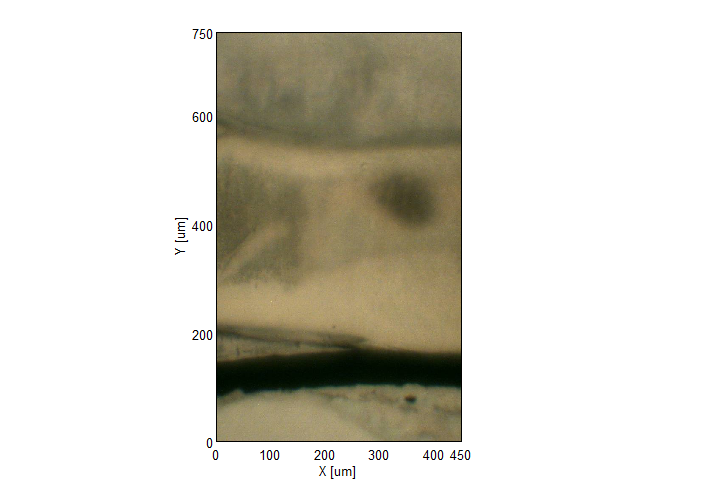
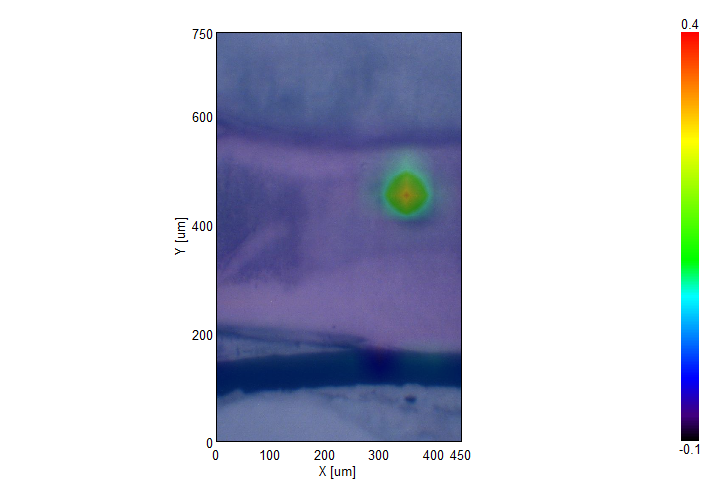
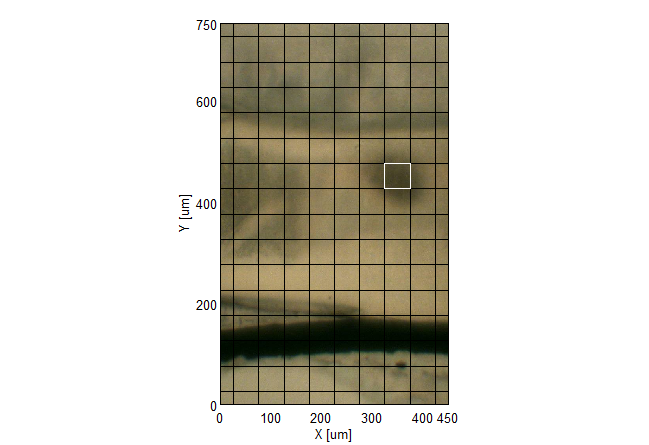
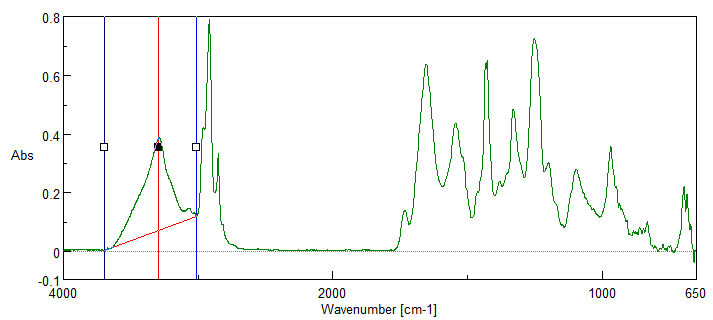
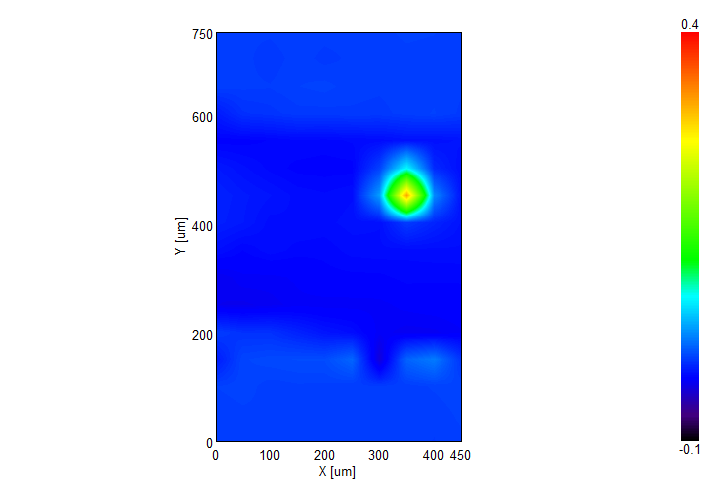
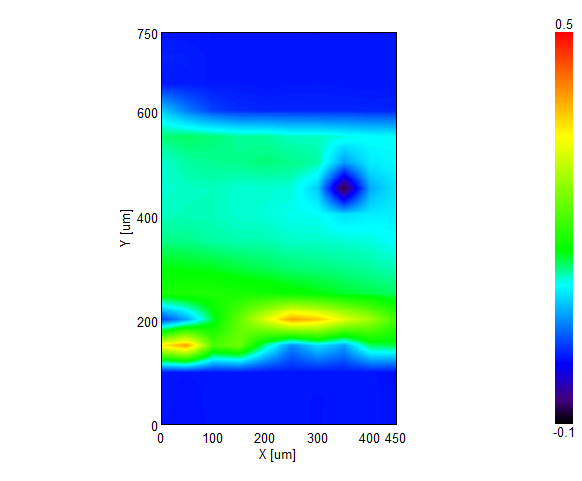
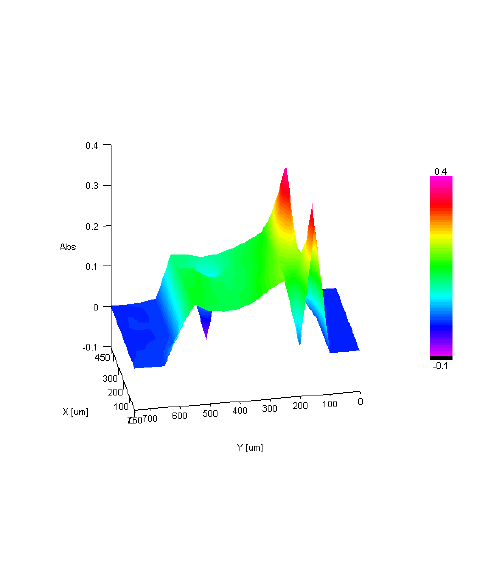
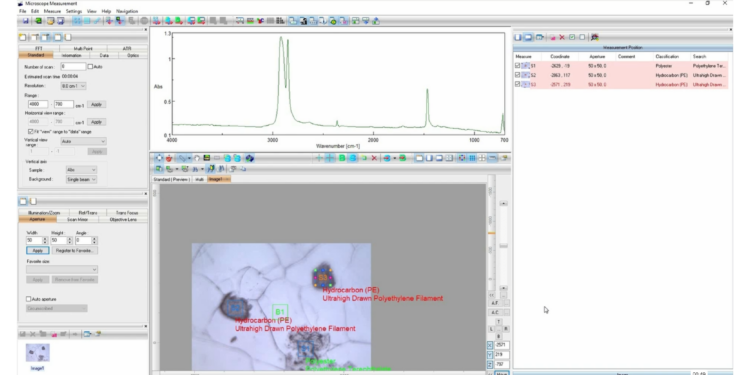
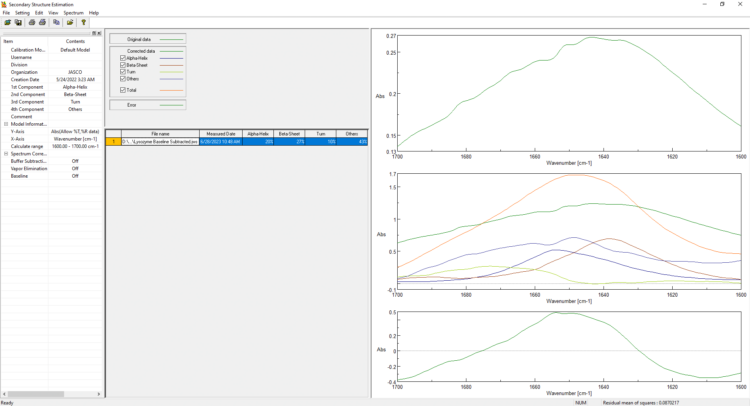
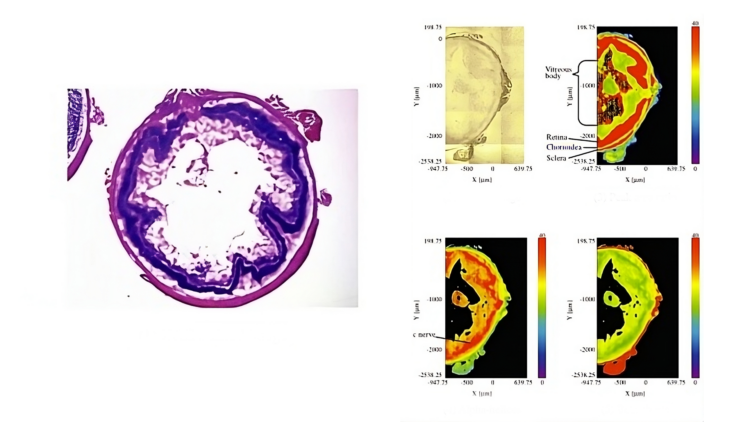
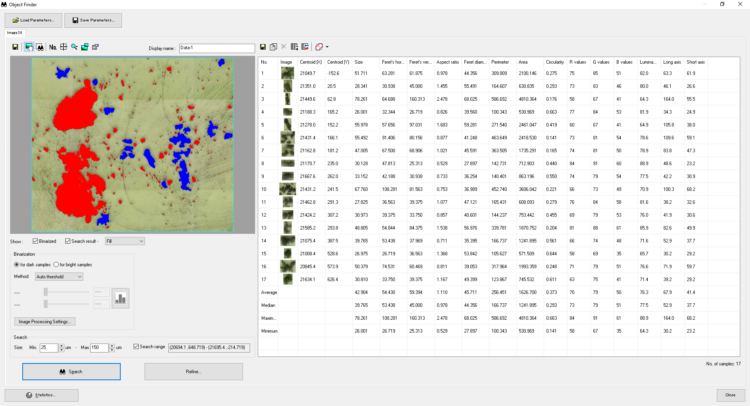
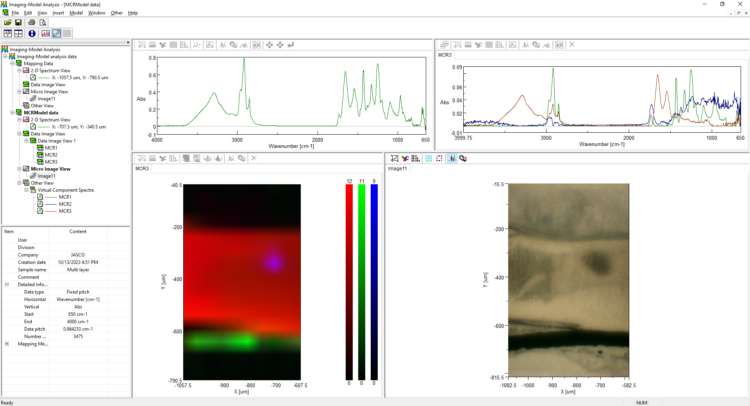
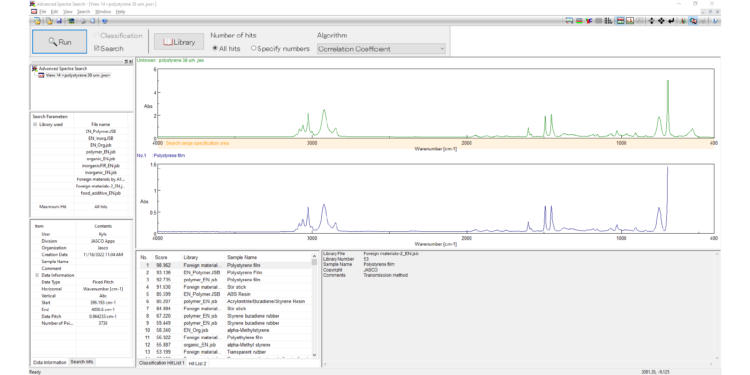
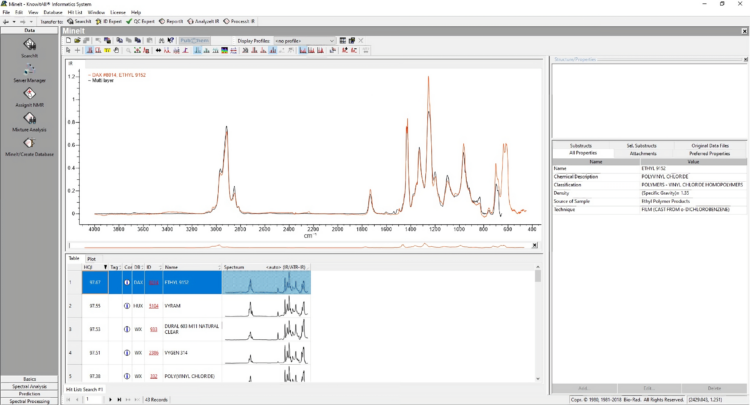
IR Microscope Imaging Analysis is used to perform post-measurement qualitative and quantitative analysis on spectral data measured obtained from the IRT- Series of FTIR microscopes; this powerful software can be used to analyze multi-data arrays from single point to complex three-dimensional measurement maps. The user interface is highly-configurable to display only the tools, data necessary for the user’s requirements – spectral data, optical image and data representations in two and three dimensions can all be displayed in different views on the screen simultaneously.